- 1 Arizona Trees
- 2 Learner classes with special methods
- 3 Resampling for comparing training on same or other subsets
- 4 Resampling for comparing train subsets and sizes
- 5 Resampling for comparing training on same or other groups
- 6 Compute resampling results in a project
- 7 Initialize a new project grid table
- 8 Combine and save results in a project
- 9 Compute several resampling jobs
- 10 Test a project with smaller data and fewer resampling iterations
- 11 P-values for comparing Same/Other/All training
- 12 P-values for full versus down-sampled SOAK results
- 13 Score benchmark results
-- A --
AZtrees
AutoTunerTorch_epochs
-- L --
LearnerRegrCVGlmnetSave
LearnerClassifCVGlmnetSave
-- P --
proj_compute()
proj_grid()
proj_results()
proj_results_save()
proj_fread()
proj_submit()
proj_compute_mpi()
proj_compute_all()
proj_todo()
proj_test()
pvalue()
pvalue_downsample()
-- R --
ResamplingSameOtherCV
ResamplingSameOtherSizesCV
ResamplingVariableSizeTrainCV
-- S --
score()
1 Arizona Trees
Description
Classification data set with polygons (groups which should not be split in CV) and subsets (region3 or region4).
Usage
data("AZtrees")Format
A data frame with 5956 observations on the following 25 variables.
region3a character vector
region4a character vector
polygona numeric vector
ya character vector
ycoordlatitude
xcoordlongitude
SAMPLE_1a numeric vector
SAMPLE_2a numeric vector
SAMPLE_3a numeric vector
SAMPLE_4a numeric vector
SAMPLE_5a numeric vector
SAMPLE_6a numeric vector
SAMPLE_7a numeric vector
SAMPLE_8a numeric vector
SAMPLE_9a numeric vector
SAMPLE_10a numeric vector
SAMPLE_11a numeric vector
SAMPLE_12a numeric vector
SAMPLE_13a numeric vector
SAMPLE_14a numeric vector
SAMPLE_15a numeric vector
SAMPLE_16a numeric vector
SAMPLE_17a numeric vector
SAMPLE_18a numeric vector
SAMPLE_19a numeric vector
SAMPLE_20a numeric vector
SAMPLE_21a numeric vector
Source
Paul Nelson Arellano, paul.arellano@nau.edu
Examples
data(AZtrees)
task.obj <- mlr3::TaskClassif$new("AZtrees3", AZtrees, target="y")
task.obj$col_roles$feature <- grep("SAMPLE", names(AZtrees), value=TRUE)
task.obj$col_roles$group <- "polygon"
task.obj$col_roles$subset <- "region3"
str(task.obj)
#> Classes 'TaskClassif', 'TaskSupervised', 'Task', 'R6' <TaskClassif:AZtrees3>
same_other_sizes_cv <- mlr3resampling::ResamplingSameOtherSizesCV$new()
same_other_sizes_cv$instantiate(task.obj)
same_other_sizes_cv$instance$iteration.dt
#> test.subset train.subsets groups test.fold
#> <char> <char> <int> <int>
#> 1: NE all 125 1
#> 2: NW all 125 1
#> 3: S all 125 1
#> 4: NE all 125 2
#> 5: NW all 125 2
#> 6: S all 125 2
#> 7: NE all 125 3
#> 8: NW all 125 3
#> 9: S all 125 3
#> 10: NE other 55 1
#> 11: NW other 104 1
#> 12: S other 91 1
#> 13: NE other 55 2
#> 14: NW other 104 2
#> 15: S other 91 2
#> 16: NE other 55 3
#> 17: NW other 104 3
#> 18: S other 91 3
#> 19: NE same 70 1
#> 20: NW same 21 1
#> 21: S same 34 1
#> 22: NE same 70 2
#> 23: NW same 21 2
#> 24: S same 34 2
#> 25: NE same 70 3
#> 26: NW same 21 3
#> 27: S same 34 3
#> test.subset train.subsets groups test.fold
#> <char> <char> <int> <int>
#> test
#> <list>
#> 1: 45,46,47,48,49,50,...[296]
#> 2: 837,838,839,840,841,842,...[376]
#> 3: 2904,2905,2906,2907,2908,2909,...[1168]
#> 4: 9,10,17,18,19,20,...[579]
#> 5: 773,774,775,776,777,778,...[877]
#> 6: 2954,2955,2956,2957,2958,2959,...[86]
#> 7: 1,2,3,4,5,6,...[589]
#> 8: 1001,1002,1003,1004,1005,1006,...[310]
#> 9: 3274,3275,4052,4053,4054,4055,...[1675]
#> 10: 45,46,47,48,49,50,...[296]
#> 11: 837,838,839,840,841,842,...[376]
#> 12: 2904,2905,2906,2907,2908,2909,...[1168]
#> 13: 9,10,17,18,19,20,...[579]
#> 14: 773,774,775,776,777,778,...[877]
#> 15: 2954,2955,2956,2957,2958,2959,...[86]
#> 16: 1,2,3,4,5,6,...[589]
#> 17: 1001,1002,1003,1004,1005,1006,...[310]
#> 18: 3274,3275,4052,4053,4054,4055,...[1675]
#> 19: 45,46,47,48,49,50,...[296]
#> 20: 837,838,839,840,841,842,...[376]
#> 21: 2904,2905,2906,2907,2908,2909,...[1168]
#> 22: 9,10,17,18,19,20,...[579]
#> 23: 773,774,775,776,777,778,...[877]
#> 24: 2954,2955,2956,2957,2958,2959,...[86]
#> 25: 1,2,3,4,5,6,...[589]
#> 26: 1001,1002,1003,1004,1005,1006,...[310]
#> 27: 3274,3275,4052,4053,4054,4055,...[1675]
#> test
#> <list>
#> train seed n.train.groups iteration
#> <list> <int> <int> <int>
#> 1: 1,2,3,4,5,6,...[4116] 1 125 1
#> 2: 1,2,3,4,5,6,...[4116] 1 125 2
#> 3: 1,2,3,4,5,6,...[4116] 1 125 3
#> 4: 1,2,3,4,5,6,...[4414] 1 125 4
#> 5: 1,2,3,4,5,6,...[4414] 1 125 5
#> 6: 1,2,3,4,5,6,...[4414] 1 125 6
#> 7: 9,10,17,18,19,20,...[3382] 1 125 7
#> 8: 9,10,17,18,19,20,...[3382] 1 125 8
#> 9: 9,10,17,18,19,20,...[3382] 1 125 9
#> 10: 773,774,775,776,777,778,...[2948] 1 55 10
#> 11: 1,2,3,4,5,6,...[2929] 1 104 11
#> 12: 1,2,3,4,5,6,...[2355] 1 91 12
#> 13: 837,838,839,840,841,842,...[3529] 1 55 13
#> 14: 1,2,3,4,5,6,...[3728] 1 104 14
#> 15: 1,2,3,4,5,6,...[1571] 1 91 15
#> 16: 773,774,775,776,777,778,...[2507] 1 55 16
#> 17: 9,10,17,18,19,20,...[2129] 1 104 17
#> 18: 9,10,17,18,19,20,...[2128] 1 91 18
#> 19: 1,2,3,4,5,6,...[1168] 1 70 19
#> 20: 773,774,775,776,777,778,...[1187] 1 21 20
#> 21: 2954,2955,2956,2957,2958,2959,...[1761] 1 34 21
#> 22: 1,2,3,4,5,6,...[885] 1 70 22
#> 23: 837,838,839,840,841,842,...[686] 1 21 23
#> 24: 2904,2905,2906,2907,2908,2909,...[2843] 1 34 24
#> 25: 9,10,17,18,19,20,...[875] 1 70 25
#> 26: 773,774,775,776,777,778,...[1253] 1 21 26
#> 27: 2904,2905,2906,2907,2908,2909,...[1254] 1 34 27
#> train seed n.train.groups iteration
#> <list> <int> <int> <int>
#> Train_subsets
#> <fctr>
#> 1: all
#> 2: all
#> 3: all
#> 4: all
#> 5: all
#> 6: all
#> 7: all
#> 8: all
#> 9: all
#> 10: other
#> 11: other
#> 12: other
#> 13: other
#> 14: other
#> 15: other
#> 16: other
#> 17: other
#> 18: other
#> 19: same
#> 20: same
#> 21: same
#> 22: same
#> 23: same
#> 24: same
#> 25: same
#> 26: same
#> 27: same
#> Train_subsets
#> <fctr>
2 Learner classes with special methods
Description
AutoTunerTorch_epochs inherits from
mlr3tuning::AutoTuner, with an initialize method that
takes arguments to construct a torch module learner. It runs gradient
descent up to max_epochs and then re-runs using the best number
of epochs. Its edit_learner method sets number of epochs to 2
(for quick learning during proj_test),
and its save_learner method returns a history data table (one
row per epoch).
LearnerRegrCVGlmnetSave inherits from
LearnerRegrCVGlmnet; its save_learner method returns a
data table of regularized linear model weights (no edit_learner
method).
LearnerClassifCVGlmnetSave is similar.
Author(s)
Toby Dylan Hocking
Examples
## Simulate regression data.
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
noise=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), each=0.5*N)))
reg.pattern.list <- list(
easy=function(x, person)x^3,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- c("x","noise")
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.task.list # two regression tasks.
#> $easy
#>
#> ── <TaskRegr> (80x3) ───────────────────────────────────────────────────────────
#> • Target: y
#> • Properties: strata
#> • Features (2):
#> • dbl (2): noise, x
#> • Strata: person
#>
#> $impossible
#>
#> ── <TaskRegr> (80x3) ───────────────────────────────────────────────────────────
#> • Target: y
#> • Properties: strata
#> • Features (2):
#> • dbl (2): noise, x
#> • Strata: person
## Create a list of learners.
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("mlr3torch") && torch::torch_is_installed()){
gen_linear <- torch::nn_module(
"my_linear",
initialize = function(task) {
self$weight = torch::nn_linear(task$n_features, 1)
},
forward = function(x) {
self$weight(x)
}
)
reg.learner.list$torch_linear <- mlr3resampling::AutoTunerTorch_epochs$new(
"torch_linear",
module_generator=gen_linear,
max_epochs=3,
batch_size=10,
loss=mlr3torch::t_loss("mse"),
measure_list=mlr3::msrs(c("regr.mse","regr.mae")))
}
if(requireNamespace("mlr3learners")){
reg.learner.list$cv_glmnet <- mlr3resampling::LearnerRegrCVGlmnetSave$new()
reg.learner.list$cv_glmnet$param_set$values$nfolds <- 3
}
reg.learner.list # a list of learners.
#> $featureless
#>
#> ── <LearnerRegrFeatureless> (regr.featureless): Featureless Regression Learner ─
#> • Model: -
#> • Parameters: robust=FALSE
#> • Packages: mlr3 and stats
#> • Predict Types: [response], se, and quantiles
#> • Feature Types: logical, integer, numeric, character, factor, ordered,
#> POSIXct, and Date
#> • Encapsulation: none (fallback: -)
#> • Properties: featureless, importance, missings, selected_features, and weights
#> • Other settings: use_weights = 'use', predict_raw = 'FALSE'
#>
#> $torch_linear
#>
#> ── <AutoTunerTorch_epochs> (torch_linear) ──────────────────────────────────────
#> • Model: -
#> • Parameters: list()
#> • Packages: mlr3, mlr3tuning, mlr3torch, and torch
#> • Predict Types: [response]
#> • Feature Types: logical, integer, numeric, character, factor, ordered,
#> POSIXct, Date, and lazy_tensor
#> • Encapsulation: none (fallback: -)
#> • Properties: featureless, hotstart_backward, hotstart_forward, importance,
#> marshal, missings, new_levels, offset, oob_error, selected_features, and
#> weights
#> • Other settings: use_weights = 'use', predict_raw = 'FALSE'
#> • Search Space:
#> id class lower upper nlevels
#> <char> <char> <num> <num> <num>
#> 1: epochs ParamInt 0 3 4
#>
#> $cv_glmnet
#>
#> ── <LearnerRegrCVGlmnetSave> (regr.cv_glmnet): GLM with Elastic Net Regularizati
#> • Model: -
#> • Parameters: family=gaussian, nfolds=3, use_pred_offset=TRUE
#> • Packages: mlr3, mlr3learners, and glmnet
#> • Predict Types: [response]
#> • Feature Types: logical, integer, and numeric
#> • Encapsulation: none (fallback: -)
#> • Properties: offset, selected_features, and weights
#> • Other settings: use_weights = 'use', predict_raw = 'FALSE'
# 2-fold CV.
kfold <- mlr3::ResamplingCV$new()
kfold$param_set$values$folds <- 2
# Create project grid.
pkg.proj.dir <- tempfile()
pgrid <- mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
score_args=mlr3::msrs("regr.rmse"),
kfold)
test_out <- mlr3resampling::proj_test(pkg.proj.dir)
test_out$learners_history.csv # from AutoTunerTorch_epochs, 2 epochs for testing.
test_out$learners_weights.csv # from LearnerRegrCVGlmnetSave
torch.job.i <- which(pgrid$learner_id=="torch_linear")[1]
mlr3resampling::proj_compute(torch.job.i, pkg.proj.dir)
mlr3resampling::proj_results_save(pkg.proj.dir)
full_out <- mlr3resampling::proj_fread(pkg.proj.dir)
full_out$learners_history.csv # from AutoTunerTorch_epochs, 3 epochs.
3 Resampling for comparing training on same or other subsets
Description
ResamplingSameOtherCV
defines how a task is partitioned for
resampling, for example in
resample() or
benchmark().
Resampling objects can be instantiated on a
Task,
which should define at least one subset variable.
After instantiation, sets can be accessed via
$train_set(i) and
$test_set(i), respectively.
Details
This provides an implementation of SOAK, Same/Other/All K-fold
cross-validation. After instantiation, this class provides information
in $instance that can be used for visualizing the
splits, as shown in the vignette. Most typical machine learning users
should instead use
ResamplingSameOtherSizesCV, which does not support these
visualization features, but provides other relevant machine learning
features, such as group role, which is not supported by
ResamplingSameOtherCV.
A supervised learning algorithm inputs a train set, and outputs a prediction function, which can be used on a test set. If each data point belongs to a subset (such as geographic region, year, etc), then how do we know if it is possible to train on one subset, and predict accurately on another subset? Cross-validation can be used to determine the extent to which this is possible, by first assigning fold IDs from 1 to K to all data (possibly using stratification, usually by subset and label). Then we loop over test sets (subset/fold combinations), train sets (same subset, other subsets, all subsets), and compute test/prediction accuracy for each combination. Comparing test/prediction accuracy between same and other, we can determine the extent to which it is possible (perfect if same/other have similar test accuracy for each subset; other is usually somewhat less accurate than same; other can be just as bad as featureless baseline when the subsets have different patterns).
Stratification
ResamplingSameOtherCV supports stratified sampling.
The stratification variables are assumed to be discrete,
and must be stored in the Task with column role "stratum".
In case of multiple stratification variables,
each combination of the values of the stratification variables forms a stratum.
Grouping
ResamplingSameOtherCV does not support grouping of
observations that should not be split in cross-validation.
See ResamplingSameOtherSizesCV for another sampler which
does support both group and subset roles.
Subsets
The subset variable is assumed to be discrete,
and must be stored in the Task with column role "subset".
The number of cross-validation folds K should be defined as the
fold parameter.
In each subset, there will be about an equal number of observations
assigned to each of the K folds.
The assignments are stored in
$instance$id.dt.
The train/test splits are defined by all possible combinations of
test subset, test fold, and train subsets (Same/Other/All).
The splits are stored in
$instance$iteration.dt.
Methods
Public methods
Method new()
Creates a new instance of this R6 class.
Usage
Resampling$new( id, param_set = ps(), duplicated_ids = FALSE, label = NA_character_, man = NA_character_ )
Arguments
id(
character(1))
Identifier for the new instance.param_set(paradox::ParamSet)
Set of hyperparameters.duplicated_ids(
logical(1))
Set toTRUEif this resampling strategy may have duplicated row ids in a single training set or test set.label(
character(1))
Label for the new instance.man(
character(1))
String in the format[pkg]::[topic]pointing to a manual page for this object. The referenced help package can be opened via method$help().
Method train_set()
Returns the row ids of the i-th training set.
Usage
Resampling$train_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
Method test_set()
Returns the row ids of the i-th test set.
Usage
Resampling$test_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
See Also
arXiv paper https://arxiv.org/abs/2410.08643 describing SOAK algorithm.
Articles https://github.com/tdhock/mlr3resampling/wiki/Articles
Package mlr3 for standard
Resampling, which does not support comparing train on Same/Other/All subsets.-
vignette(package="mlr3resampling")for more detailed examples.
Examples
same_other <- mlr3resampling::ResamplingSameOtherCV$new()
same_other$param_set$values$folds <- 5
4 Resampling for comparing train subsets and sizes
Description
ResamplingSameOtherSizesCV
defines how a task is partitioned for
resampling, for example in
resample() or
benchmark().
Resampling objects can be instantiated on a
Task,
which can use the subset role.
After instantiation, sets can be accessed via
$train_set(i) and
$test_set(i), respectively.
Details
This is an implementation of SOAK, Same/Other/All K-fold cross-validation. A supervised learning algorithm inputs a train set, and outputs a prediction function, which can be used on a test set. If each data point belongs to a subset (such as geographic region, year, etc), then how do we know if it is possible to train on one subset, and predict accurately on another subset? Cross-validation can be used to determine the extent to which this is possible, by first assigning fold IDs from 1 to K to all data (possibly using stratification, usually by subset and label). Then we loop over test sets (subset/fold combinations), train sets (same subset, other subsets, all subsets), and compute test/prediction accuracy for each combination. Comparing test/prediction accuracy between same and other, we can determine the extent to which it is possible (perfect if same/other have similar test accuracy for each subset; other is usually somewhat less accurate than same; other can be just as bad as featureless baseline when the subsets have different patterns).
This class has more parameters/potential applications than
ResamplingSameOtherCV and
ResamplingVariableSizeTrainCV,
which are older and should only be preferred
for visualization purposes.
Stratification
ResamplingSameOtherSizesCV supports stratified sampling.
The stratification variables are assumed to be discrete,
and must be stored in the Task with column role "stratum".
In case of multiple stratification variables,
each combination of the values of the stratification variables forms a stratum.
Grouping
ResamplingSameOtherSizesCV supports grouping of
observations that will not be split in cross-validation.
The grouping variable is assumed to be discrete,
and must be stored in the Task with column role
"group".
Subsets
ResamplingSameOtherSizesCV supports training on different
subsets of observations.
The subset variable is assumed to be discrete,
and must be stored in the Task with column role "subset".
Parameters
The number of cross-validation folds K should be defined as the
fold parameter, default 3.
The number of random seeds for down-sampling should be defined as the
seeds parameter, default 1.
The ratio for down-sampling should be defined as the ratio
parameter, default 0.5. The min size of same and other sets is
repeatedly multiplied by this ratio, to obtain smaller sample sizes.
The number of down-sampling sizes/multiplications should be defined as
the sizes parameter, which can also take two special values:
default -1 means no down-sampling at all, and 0 means only down-sampling
to the sizes of the same/other sets.
The ignore_subset parameter should be either TRUE or
FALSE (default), whether to ignore the subset
role. TRUE only creates splits for same subset (even if task
defines subset role), and is useful for subtrain/validation
splits (hyper-parameter learning). Note that this feature will work on a
task with both stratum and group roles (unlike
ResamplingCV).
The subsets parameter should specify the train subsets of
interest: "S" (same),
"O" (other), "A" (all), "SO", "SA",
"SOA" (default).
In each subset, there will be about an equal number of observations
assigned to each of the K folds.
The train/test splits are defined by all possible combinations of
test subset, test fold, train subsets (same/other/all), down-sampling
sizes, and random seeds.
The splits are stored in
$instance$iteration.dt.
Methods
Public methods
Method new()
Creates a new instance of this R6 class.
Usage
Resampling$new( id, param_set = ps(), duplicated_ids = FALSE, label = NA_character_, man = NA_character_ )
Arguments
id(
character(1))
Identifier for the new instance.param_set(paradox::ParamSet)
Set of hyperparameters.duplicated_ids(
logical(1))
Set toTRUEif this resampling strategy may have duplicated row ids in a single training set or test set.label(
character(1))
Label for the new instance.man(
character(1))
String in the format[pkg]::[topic]pointing to a manual page for this object. The referenced help package can be opened via method$help().
Method train_set()
Returns the row ids of the i-th training set.
Usage
Resampling$train_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
Method test_set()
Returns the row ids of the i-th test set.
Usage
Resampling$test_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
See Also
arXiv paper https://arxiv.org/abs/2410.08643 describing SOAK algorithm.
Articles https://github.com/tdhock/mlr3resampling/wiki/Articles
Package mlr3 for standard
Resampling, which does not support comparing train on Same/Other/All subsets.-
vignette(package="mlr3resampling")for more detailed examples.
Examples
same_other_sizes <- mlr3resampling::ResamplingSameOtherSizesCV$new()
same_other_sizes$param_set$values$folds <- 5
5 Resampling for comparing training on same or other groups
Description
ResamplingVariableSizeTrainCV
defines how a task is partitioned for
resampling, for example in
resample() or
benchmark().
Resampling objects can be instantiated on a
Task.
After instantiation, sets can be accessed via
$train_set(i) and
$test_set(i), respectively.
Details
A supervised learning algorithm inputs a train set, and outputs a prediction function, which can be used on a test set. How many train samples are required to get accurate predictions on a test set? Cross-validation can be used to answer this question, with variable size train sets.
Stratification
ResamplingVariableSizeTrainCV supports stratified sampling.
The stratification variables are assumed to be discrete,
and must be stored in the Task with column role "stratum".
In case of multiple stratification variables,
each combination of the values of the stratification variables forms a stratum.
Grouping
ResamplingVariableSizeTrainCV
does not support grouping of observations.
Hyper-parameters
The number of cross-validation folds should be defined as the
fold parameter.
For each fold ID, the corresponding observations are considered the test set, and a variable number of other observations are considered the train set.
The random_seeds parameter controls the number of random
orderings of the train set that are considered.
For each random order of the train set, the min_train_data
parameter controls the size of the smallest stratum in the smallest
train set considered.
To determine the other train set sizes, we use an equally spaced grid
on the log scale, from min_train_data to the largest train set
size (all data not in test set). The
number of train set sizes in this grid is determined by the
train_sizes parameter.
Methods
Public methods
Method new()
Creates a new instance of this R6 class.
Usage
Resampling$new( id, param_set = ps(), duplicated_ids = FALSE, label = NA_character_, man = NA_character_ )
Arguments
id(
character(1))
Identifier for the new instance.param_set(paradox::ParamSet)
Set of hyperparameters.duplicated_ids(
logical(1))
Set toTRUEif this resampling strategy may have duplicated row ids in a single training set or test set.label(
character(1))
Label for the new instance.man(
character(1))
String in the format[pkg]::[topic]pointing to a manual page for this object. The referenced help package can be opened via method$help().
Method train_set()
Returns the row ids of the i-th training set.
Usage
Resampling$train_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
Method test_set()
Returns the row ids of the i-th test set.
Usage
Resampling$test_set(i)
Arguments
i(
integer(1))
Iteration.
Returns
(integer()) of row ids.
Examples
(var_sizes <- mlr3resampling::ResamplingVariableSizeTrainCV$new())
#> ── <ResamplingVariableSizeTrainCV> : Cross-Validation with variable size train s
#> • Iterations:
#> • Instantiated: FALSE
#> • Parameters: folds=3, min_train_data=10, random_seeds=3, train_sizes=5
6 Compute resampling results in a project
Description
Runs train() and predict(), then a data table
with one row is saved to an RDS file in the grid_jobs
directory.
Usage
proj_compute(grid_job_i, proj_dir, verbose=FALSE, process_fun=Sys.getpid)Arguments
grid_job_i |
integer from 1 to number of jobs (rows in |
proj_dir |
Project directory created by |
verbose |
Logical: print messages? |
process_fun |
function called with no arguments
(default |
Details
If everything goes well, the user should not need to run this
function.
Instead, the user runs proj_submit as Step 2 out of the
typical 3 step pipeline (init grid, submit, read results).
proj_compute can sometimes be useful for testing or debugging the submit step,
since it runs one split at a time.
Value
proj_compute returns a data table with one row of results.
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), each=0.5*N)))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- "x"
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
pkg.proj.dir <- tempfile()
mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
SOAK,
order_jobs = function(DT)1:2, # for CRAN.
score_args=mlr3::msrs(c("regr.rmse", "regr.mae")))
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> 2: 1 1 1 easy regr.featureless same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <fctr> <char> <int> <int> <int> <int> <int>
#> 1: Alice all 52 1 1 52 1
#> 2: Bob all 52 1 1 52 2
#> Train_subsets
#> <fctr>
#> 1: all
#> 2: all
mlr3resampling::proj_compute(1, pkg.proj.dir)
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <char> <char> <int> <int> <int> <int> <int>
#> 1: Alice all 52 1 1 52 1
#> Train_subsets start.time end.time process learner pred
#> <char> <POSc> <POSc> <int> <list> <list>
#> 1: all 2026-04-28 14:59:28 2026-04-28 14:59:28 26324 [NULL] [NULL]
#> regr.rmse regr.mae
#> <num> <num>
#> 1: 1.239123 0.86461
7 Initialize a new project grid table
Description
A project grid consists of all combinations of tasks, learners, resampling types, and resampling iterations, to be computed in parallel. This function creates a project directory with files to describe the grid.
Usage
proj_grid(
proj_dir, tasks, learners, resamplings,
order_jobs = NULL, score_args = NULL,
save_learner = save_learner_default, save_pred = FALSE,
train_seed = 1L, resampling_seed = 1L)Arguments
proj_dir |
Path to directory to create. |
tasks |
List of Tasks, or a single Task. |
learners |
List of Learners, or a single Learner. |
resamplings |
List of Resamplings, or a single Resampling. |
order_jobs |
Function which takes split table as input, and
returns integer vector of row numbers of the split table to write to
|
score_args |
Passed to |
save_learner |
Function to process Learner, after
training/prediction, but before saving result to disk.
For interpreting complex models, you should write a function that
returns only the parts of the model that you need (and discards the
other parts which would take up disk space for no reason).
Default is to call |
save_pred |
Function to process Prediction before saving to disk.
Default |
train_seed |
integer: random seed to set before training (default
|
resampling_seed |
integer: random seed to set before
instantiating each resampling (default |
Details
This is Step 1 out of the
typical 3 step pipeline (init grid, submit, read results).
It creates a grid_jobs.csv table which has a column status;
each row is initialized to "not started" or "done",
depending on whether the corresponding result RDS file exists already.
Value
Data table of splits to be processed (same as table saved to grid_jobs.rds).
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), each=0.5*N)))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- "x"
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
pkg.proj.dir <- tempfile()
mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
SOAK,
score_args=mlr3::msrs(c("regr.rmse", "regr.mae")))
#> task.i learner.i resampling.i task_id learner_id
#> <int> <int> <int> <char> <char>
#> 1: 1 1 1 easy regr.featureless
#> 2: 1 1 1 easy regr.featureless
#> 3: 1 1 1 easy regr.featureless
#> 4: 1 1 1 easy regr.featureless
#> 5: 1 1 1 easy regr.featureless
#> 6: 1 1 1 easy regr.featureless
#> 7: 1 2 1 easy regr.rpart
#> 8: 1 2 1 easy regr.rpart
#> 9: 1 2 1 easy regr.rpart
#> 10: 1 2 1 easy regr.rpart
#> 11: 1 2 1 easy regr.rpart
#> 12: 1 2 1 easy regr.rpart
#> 13: 2 1 1 impossible regr.featureless
#> 14: 2 1 1 impossible regr.featureless
#> 15: 2 1 1 impossible regr.featureless
#> 16: 2 1 1 impossible regr.featureless
#> 17: 2 1 1 impossible regr.featureless
#> 18: 2 1 1 impossible regr.featureless
#> 19: 2 2 1 impossible regr.rpart
#> 20: 2 2 1 impossible regr.rpart
#> 21: 2 2 1 impossible regr.rpart
#> 22: 2 2 1 impossible regr.rpart
#> 23: 2 2 1 impossible regr.rpart
#> 24: 2 2 1 impossible regr.rpart
#> 25: 1 1 1 easy regr.featureless
#> 26: 1 1 1 easy regr.featureless
#> 27: 1 1 1 easy regr.featureless
#> 28: 1 1 1 easy regr.featureless
#> 29: 1 1 1 easy regr.featureless
#> 30: 1 1 1 easy regr.featureless
#> 31: 1 1 1 easy regr.featureless
#> 32: 1 1 1 easy regr.featureless
#> 33: 1 1 1 easy regr.featureless
#> 34: 1 1 1 easy regr.featureless
#> 35: 1 1 1 easy regr.featureless
#> 36: 1 1 1 easy regr.featureless
#> 37: 1 2 1 easy regr.rpart
#> 38: 1 2 1 easy regr.rpart
#> 39: 1 2 1 easy regr.rpart
#> 40: 1 2 1 easy regr.rpart
#> 41: 1 2 1 easy regr.rpart
#> 42: 1 2 1 easy regr.rpart
#> 43: 1 2 1 easy regr.rpart
#> 44: 1 2 1 easy regr.rpart
#> 45: 1 2 1 easy regr.rpart
#> 46: 1 2 1 easy regr.rpart
#> 47: 1 2 1 easy regr.rpart
#> 48: 1 2 1 easy regr.rpart
#> 49: 2 1 1 impossible regr.featureless
#> 50: 2 1 1 impossible regr.featureless
#> 51: 2 1 1 impossible regr.featureless
#> 52: 2 1 1 impossible regr.featureless
#> 53: 2 1 1 impossible regr.featureless
#> 54: 2 1 1 impossible regr.featureless
#> 55: 2 1 1 impossible regr.featureless
#> 56: 2 1 1 impossible regr.featureless
#> 57: 2 1 1 impossible regr.featureless
#> 58: 2 1 1 impossible regr.featureless
#> 59: 2 1 1 impossible regr.featureless
#> 60: 2 1 1 impossible regr.featureless
#> 61: 2 2 1 impossible regr.rpart
#> 62: 2 2 1 impossible regr.rpart
#> 63: 2 2 1 impossible regr.rpart
#> 64: 2 2 1 impossible regr.rpart
#> 65: 2 2 1 impossible regr.rpart
#> 66: 2 2 1 impossible regr.rpart
#> 67: 2 2 1 impossible regr.rpart
#> 68: 2 2 1 impossible regr.rpart
#> 69: 2 2 1 impossible regr.rpart
#> 70: 2 2 1 impossible regr.rpart
#> 71: 2 2 1 impossible regr.rpart
#> 72: 2 2 1 impossible regr.rpart
#> task.i learner.i resampling.i task_id learner_id
#> <int> <int> <int> <char> <char>
#> resampling_id test.subset train.subsets groups test.fold seed
#> <char> <fctr> <char> <int> <int> <int>
#> 1: same_other_sizes_cv Alice all 52 1 1
#> 2: same_other_sizes_cv Bob all 52 1 1
#> 3: same_other_sizes_cv Alice all 52 2 1
#> 4: same_other_sizes_cv Bob all 52 2 1
#> 5: same_other_sizes_cv Alice all 52 3 1
#> 6: same_other_sizes_cv Bob all 52 3 1
#> 7: same_other_sizes_cv Alice all 52 1 1
#> 8: same_other_sizes_cv Bob all 52 1 1
#> 9: same_other_sizes_cv Alice all 52 2 1
#> 10: same_other_sizes_cv Bob all 52 2 1
#> 11: same_other_sizes_cv Alice all 52 3 1
#> 12: same_other_sizes_cv Bob all 52 3 1
#> 13: same_other_sizes_cv Alice all 52 1 1
#> 14: same_other_sizes_cv Bob all 52 1 1
#> 15: same_other_sizes_cv Alice all 52 2 1
#> 16: same_other_sizes_cv Bob all 52 2 1
#> 17: same_other_sizes_cv Alice all 52 3 1
#> 18: same_other_sizes_cv Bob all 52 3 1
#> 19: same_other_sizes_cv Alice all 52 1 1
#> 20: same_other_sizes_cv Bob all 52 1 1
#> 21: same_other_sizes_cv Alice all 52 2 1
#> 22: same_other_sizes_cv Bob all 52 2 1
#> 23: same_other_sizes_cv Alice all 52 3 1
#> 24: same_other_sizes_cv Bob all 52 3 1
#> 25: same_other_sizes_cv Alice other 26 1 1
#> 26: same_other_sizes_cv Bob other 26 1 1
#> 27: same_other_sizes_cv Alice other 26 2 1
#> 28: same_other_sizes_cv Bob other 26 2 1
#> 29: same_other_sizes_cv Alice other 26 3 1
#> 30: same_other_sizes_cv Bob other 26 3 1
#> 31: same_other_sizes_cv Alice same 26 1 1
#> 32: same_other_sizes_cv Bob same 26 1 1
#> 33: same_other_sizes_cv Alice same 26 2 1
#> 34: same_other_sizes_cv Bob same 26 2 1
#> 35: same_other_sizes_cv Alice same 26 3 1
#> 36: same_other_sizes_cv Bob same 26 3 1
#> 37: same_other_sizes_cv Alice other 26 1 1
#> 38: same_other_sizes_cv Bob other 26 1 1
#> 39: same_other_sizes_cv Alice other 26 2 1
#> 40: same_other_sizes_cv Bob other 26 2 1
#> 41: same_other_sizes_cv Alice other 26 3 1
#> 42: same_other_sizes_cv Bob other 26 3 1
#> 43: same_other_sizes_cv Alice same 26 1 1
#> 44: same_other_sizes_cv Bob same 26 1 1
#> 45: same_other_sizes_cv Alice same 26 2 1
#> 46: same_other_sizes_cv Bob same 26 2 1
#> 47: same_other_sizes_cv Alice same 26 3 1
#> 48: same_other_sizes_cv Bob same 26 3 1
#> 49: same_other_sizes_cv Alice other 26 1 1
#> 50: same_other_sizes_cv Bob other 26 1 1
#> 51: same_other_sizes_cv Alice other 26 2 1
#> 52: same_other_sizes_cv Bob other 26 2 1
#> 53: same_other_sizes_cv Alice other 26 3 1
#> 54: same_other_sizes_cv Bob other 26 3 1
#> 55: same_other_sizes_cv Alice same 26 1 1
#> 56: same_other_sizes_cv Bob same 26 1 1
#> 57: same_other_sizes_cv Alice same 26 2 1
#> 58: same_other_sizes_cv Bob same 26 2 1
#> 59: same_other_sizes_cv Alice same 26 3 1
#> 60: same_other_sizes_cv Bob same 26 3 1
#> 61: same_other_sizes_cv Alice other 26 1 1
#> 62: same_other_sizes_cv Bob other 26 1 1
#> 63: same_other_sizes_cv Alice other 26 2 1
#> 64: same_other_sizes_cv Bob other 26 2 1
#> 65: same_other_sizes_cv Alice other 26 3 1
#> 66: same_other_sizes_cv Bob other 26 3 1
#> 67: same_other_sizes_cv Alice same 26 1 1
#> 68: same_other_sizes_cv Bob same 26 1 1
#> 69: same_other_sizes_cv Alice same 26 2 1
#> 70: same_other_sizes_cv Bob same 26 2 1
#> 71: same_other_sizes_cv Alice same 26 3 1
#> 72: same_other_sizes_cv Bob same 26 3 1
#> resampling_id test.subset train.subsets groups test.fold seed
#> <char> <fctr> <char> <int> <int> <int>
#> n.train.groups iteration Train_subsets
#> <int> <int> <fctr>
#> 1: 52 1 all
#> 2: 52 2 all
#> 3: 52 3 all
#> 4: 52 4 all
#> 5: 52 5 all
#> 6: 52 6 all
#> 7: 52 1 all
#> 8: 52 2 all
#> 9: 52 3 all
#> 10: 52 4 all
#> 11: 52 5 all
#> 12: 52 6 all
#> 13: 52 1 all
#> 14: 52 2 all
#> 15: 52 3 all
#> 16: 52 4 all
#> 17: 52 5 all
#> 18: 52 6 all
#> 19: 52 1 all
#> 20: 52 2 all
#> 21: 52 3 all
#> 22: 52 4 all
#> 23: 52 5 all
#> 24: 52 6 all
#> 25: 26 7 other
#> 26: 26 8 other
#> 27: 26 9 other
#> 28: 26 10 other
#> 29: 26 11 other
#> 30: 26 12 other
#> 31: 26 13 same
#> 32: 26 14 same
#> 33: 26 15 same
#> 34: 26 16 same
#> 35: 26 17 same
#> 36: 26 18 same
#> 37: 26 7 other
#> 38: 26 8 other
#> 39: 26 9 other
#> 40: 26 10 other
#> 41: 26 11 other
#> 42: 26 12 other
#> 43: 26 13 same
#> 44: 26 14 same
#> 45: 26 15 same
#> 46: 26 16 same
#> 47: 26 17 same
#> 48: 26 18 same
#> 49: 26 7 other
#> 50: 26 8 other
#> 51: 26 9 other
#> 52: 26 10 other
#> 53: 26 11 other
#> 54: 26 12 other
#> 55: 26 13 same
#> 56: 26 14 same
#> 57: 26 15 same
#> 58: 26 16 same
#> 59: 26 17 same
#> 60: 26 18 same
#> 61: 26 7 other
#> 62: 26 8 other
#> 63: 26 9 other
#> 64: 26 10 other
#> 65: 26 11 other
#> 66: 26 12 other
#> 67: 26 13 same
#> 68: 26 14 same
#> 69: 26 15 same
#> 70: 26 16 same
#> 71: 26 17 same
#> 72: 26 18 same
#> n.train.groups iteration Train_subsets
#> <int> <int> <fctr>
mlr3resampling::proj_compute(2, pkg.proj.dir)
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <char> <char> <int> <int> <int> <int> <int>
#> 1: Bob all 52 1 1 52 2
#> Train_subsets start.time end.time process learner pred
#> <char> <POSc> <POSc> <int> <list> <list>
#> 1: all 2026-04-28 14:59:28 2026-04-28 14:59:28 26324 [NULL] [NULL]
#> regr.rmse regr.mae
#> <num> <num>
#> 1: 1.215372 1.030797
8 Combine and save results in a project
Description
proj_results globs the RDS result files in the project
directory, and combines them into a result table via rbindlist().
proj_results_save saves that result table to results.rds
and one or more CSV files (redundant with RDS data, but more
convenient).
proj_fread reads the CSV files, adding columns from
proj_grid.csv to learners*.csv.
Usage
proj_results(proj_dir, verbose=FALSE)
proj_results_save(proj_dir, verbose=FALSE)
proj_fread(proj_dir)Arguments
proj_dir |
Project directory created via
|
verbose |
logical: cat progress? (default FALSE) |
Details
This is Step 3 out of the typical 3 step pipeline (init grid, submit, read results).
Actually, if step 2 worked as intended, then
proj_results_save is called at the end of step 2,
which saves result files to disk that you can read directly:
results.csvcontains test measures for each split.
results.rdscontains additional list columns for
learnerandpred(useful for model interpretation), and can be read viareadRDS()learners.csvonly exists if
learnercolumn is a data frame, in which case it contains the atomic columns, along with meta-data describing each split.learners_*.csvonly exists if
learnercolumn is a named list of data frames: star in file name is expanded using list names, with CSV data taken from atomic columns.
Value
proj_results returns a data table with all columns, whereas
proj_results_save returns the same table with only atomic columns.
proj_fread returns a list
with names corresponding to CSV files in the test directory, and
values are the data tables that result from fread.
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
noise=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), each=0.5*N)))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- c("x","noise")
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
pkg.proj.dir <- tempfile()
mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
SOAK,
save_learner=function(L){
if(inherits(L, "LearnerRegrRpart")){
list(rpart=L$model$frame)
}
},
order_jobs = function(DT)which(DT$iteration==1)[1:2], # for CRAN.
score_args=mlr3::msrs(c("regr.rmse", "regr.mae")))
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> 2: 1 2 1 easy regr.rpart same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <fctr> <char> <int> <int> <int> <int> <int>
#> 1: Alice all 52 1 1 52 1
#> 2: Alice all 52 1 1 52 1
#> Train_subsets
#> <fctr>
#> 1: all
#> 2: all
computed <- mlr3resampling::proj_compute_all(pkg.proj.dir)
result_list <- mlr3resampling::proj_fread(pkg.proj.dir)
result_list$learners_rpart.csv # one row per node in decision tree.
#> grid_job_i var n wt dev yval complexity ncompete
#> <int> <char> <int> <int> <num> <num> <num> <int>
#> 1: 2 x 52 52 72.962344 1.0985339 0.340563892 1
#> 2: 2 x 45 45 46.090929 0.8479447 0.340563892 1
#> 3: 2 <leaf> 34 34 8.126498 0.3936568 0.008652142 0
#> 4: 2 <leaf> 11 11 9.259200 2.2521073 0.010000000 0
#> 5: 2 <leaf> 7 7 5.879966 2.7094642 0.010000000 0
#> nsurrogate task_id learner_id resampling_id test.subset train.subsets
#> <int> <char> <char> <char> <char> <char>
#> 1: 1 easy regr.rpart same_other_sizes_cv Alice all
#> 2: 0 easy regr.rpart same_other_sizes_cv Alice all
#> 3: 0 easy regr.rpart same_other_sizes_cv Alice all
#> 4: 0 easy regr.rpart same_other_sizes_cv Alice all
#> 5: 0 easy regr.rpart same_other_sizes_cv Alice all
#> groups test.fold seed n.train.groups iteration Train_subsets
#> <int> <int> <int> <int> <int> <char>
#> 1: 52 1 1 52 1 all
#> 2: 52 1 1 52 1 all
#> 3: 52 1 1 52 1 all
#> 4: 52 1 1 52 1 all
#> 5: 52 1 1 52 1 all
result_list$results.csv # test error in regr.* columns.
#> grid_job_i task.i learner.i resampling.i task_id learner_id
#> <int> <int> <int> <int> <char> <char>
#> 1: 1 1 1 1 easy regr.featureless
#> 2: 2 1 2 1 easy regr.rpart
#> resampling_id test.subset train.subsets groups test.fold seed
#> <char> <char> <char> <int> <int> <int>
#> 1: same_other_sizes_cv Alice all 52 1 1
#> 2: same_other_sizes_cv Alice all 52 1 1
#> n.train.groups iteration Train_subsets start.time
#> <int> <int> <char> <POSc>
#> 1: 52 1 all 2026-04-28 14:59:29
#> 2: 52 1 all 2026-04-28 14:59:29
#> end.time process regr.rmse regr.mae
#> <POSc> <int> <num> <num>
#> 1: 2026-04-28 14:59:29 26324 1.0207548 0.7546977
#> 2: 2026-04-28 14:59:29 26324 0.7706187 0.6652567
9 Compute several resampling jobs
Description
proj_todo determines which jobs remain to be computed.
proj_compute_all computes all remaining jobs using
future_lapply if available, otherwise
lapply.
proj_compute_mpi computes all remaining jobs in parallel using
MPI (should be run in an R session activated by mpirun or
srun).
proj_submit is a non-blocking call to SLURM sbatch,
asking for a single job with several tasks that run proj_compute_mpi.
Usage
proj_todo(proj_dir)
proj_compute_mpi(proj_dir, verbose=FALSE)
proj_compute_all(proj_dir, verbose=FALSE, LAPPLY)
proj_submit(
proj_dir, tasks = 2, hours = 1, gigabytes = 1,
verbose = FALSE)Arguments
proj_dir |
Project directory created via |
tasks |
Positive integer: |
hours |
Hours of walltime to ask the SLURM scheduler. |
gigabytes |
Gigabytes of memory to ask the SLURM scheduler. |
verbose |
Logical: print messages? |
LAPPLY |
Function like |
Details
This is Step 2 out of the typical 3 step pipeline (init grid, submit, read results).
Value
proj_submit returns the ID of the submitted SLURM job.
proj_compute_all and proj_compute_mpi return a data
table of results computed.
proj_todo returns a vector of job IDs not yet computed.
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), each=0.5*N)))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- "x"
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
pkg.proj.dir <- tempfile()
mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
SOAK,
order_jobs = function(DT)1:2, # for CRAN.
score_args=mlr3::msrs(c("regr.rmse", "regr.mae")))
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> 2: 1 1 1 easy regr.featureless same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <fctr> <char> <int> <int> <int> <int> <int>
#> 1: Alice all 52 1 1 52 1
#> 2: Bob all 52 1 1 52 2
#> Train_subsets
#> <fctr>
#> 1: all
#> 2: all
if(requireNamespace("future.apply"))future::plan("multisession")
mlr3resampling::proj_compute_all(pkg.proj.dir)
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless same_other_sizes_cv
#> 2: 1 1 1 easy regr.featureless same_other_sizes_cv
#> test.subset train.subsets groups test.fold seed n.train.groups iteration
#> <char> <char> <int> <int> <int> <int> <int>
#> 1: Alice all 52 1 1 52 1
#> 2: Bob all 52 1 1 52 2
#> Train_subsets start.time end.time process learner pred
#> <char> <POSc> <POSc> <int> <list> <list>
#> 1: all 2026-04-28 14:59:34 2026-04-28 14:59:34 26407 [NULL] [NULL]
#> 2: all 2026-04-28 14:59:37 2026-04-28 14:59:37 26410 [NULL] [NULL]
#> regr.rmse regr.mae
#> <num> <num>
#> 1: 1.239123 0.864610
#> 2: 1.215372 1.030797
if(requireNamespace("future.apply"))future::plan("sequential")
10 Test a project with smaller data and fewer resampling iterations
Description
Like testJob,
"test" means trying an example with a few small jobs
(default one train/test split per algorithm and data set)
before running the big calculation with all jobs.
Runs proj_grid to create a new project in the
test sub-directory, with a smaller
number of samples in each task, and with only one iteration
per Resampling. Runs proj_compute_all on this new
test project, and then reads any CSV result files.
Usage
proj_test(
proj_dir, min_samples_per_stratum = 10,
edit_learner=edit_learner_default, max_jobs=Inf,
verbose=FALSE, LAPPLY=NULL)Arguments
proj_dir |
Project directory created by |
min_samples_per_stratum |
Minimum number of samples to include in the smallest stratum. Other strata will be down-sampled proportionally. |
edit_learner |
Function which inputs a learner object, and
changes it to take less time for testing. Default calls
|
max_jobs |
Numeric, max number of jobs to test (default Inf). |
verbose |
Logical: print messages? |
LAPPLY |
Function like |
Value
Same value as proj_fread on test project (list of data tables).
Author(s)
Toby Dylan Hocking
Examples
library(data.table)
N <- 8000
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=factor(rep(c("Alice","Bob"), c(0.1,0.9)*N)))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^as.integer(person))
kfold <- mlr3::ResamplingCV$new()
kfold$param_set$values$folds <- 2
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
task.dt <- data.table(reg.dt)[
, y := f(x,person)+rnorm(N, sd=0.5)
][]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target="y")
task.obj$col_roles$feature <- "x"
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
featureless=mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
pkg.proj.dir <- tempfile()
mlr3resampling::proj_grid(
pkg.proj.dir,
reg.task.list,
reg.learner.list,
kfold,
save_learner=function(L){
if(inherits(L, "LearnerRegrRpart")){
list(rpart=L$model$frame)
}
},
score_args=mlr3::msrs(c("regr.rmse", "regr.mae")))
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless cv
#> 2: 1 1 1 easy regr.featureless cv
#> 3: 1 2 1 easy regr.rpart cv
#> 4: 1 2 1 easy regr.rpart cv
#> 5: 2 1 1 impossible regr.featureless cv
#> 6: 2 1 1 impossible regr.featureless cv
#> 7: 2 2 1 impossible regr.rpart cv
#> 8: 2 2 1 impossible regr.rpart cv
#> iteration
#> <int>
#> 1: 1
#> 2: 2
#> 3: 1
#> 4: 2
#> 5: 1
#> 6: 2
#> 7: 1
#> 8: 2
mlr3resampling::proj_test(pkg.proj.dir)
#> $grid_jobs.csv
#> task.i learner.i resampling.i task_id learner_id resampling_id
#> <int> <int> <int> <char> <char> <char>
#> 1: 1 1 1 easy regr.featureless cv
#> 2: 1 2 1 easy regr.rpart cv
#> 3: 2 1 1 impossible regr.featureless cv
#> 4: 2 2 1 impossible regr.rpart cv
#> iteration
#> <int>
#> 1: 1
#> 2: 1
#> 3: 1
#> 4: 1
#>
#> $learners_rpart.csv
#> grid_job_i var n wt dev yval complexity ncompete
#> <int> <char> <int> <int> <num> <num> <num> <int>
#> 1: 2 x 50 50 78.058677 1.3852666 0.390770203 0
#> 2: 2 x 42 42 49.421715 1.0680160 0.390770203 0
#> 3: 2 x 33 33 10.448144 0.5941131 0.023122016 0
#> 4: 2 x 26 26 7.218463 0.4727662 0.018194344 0
#> 5: 2 <leaf> 14 14 2.731358 0.2563854 0.010000000 0
#> 6: 2 <leaf> 12 12 3.066878 0.7252104 0.010000000 0
#> 7: 2 <leaf> 7 7 1.424807 1.0448303 0.010000000 0
#> 8: 2 <leaf> 9 9 4.387637 2.8056601 0.010000000 0
#> 9: 2 <leaf> 8 8 2.216886 3.0508319 0.010000000 0
#> 10: 4 x 50 50 111.331741 1.1434664 0.249241891 0
#> 11: 4 x 39 39 74.483871 0.7478274 0.182219184 0
#> 12: 4 <leaf> 31 31 11.676839 0.3814418 0.005709912 0
#> 13: 4 <leaf> 8 8 42.520253 2.1675716 0.010000000 0
#> 14: 4 <leaf> 11 11 9.099336 2.5461863 0.010000000 0
#> nsurrogate task_id learner_id resampling_id iteration
#> <int> <char> <char> <char> <int>
#> 1: 0 easy regr.rpart cv 1
#> 2: 0 easy regr.rpart cv 1
#> 3: 0 easy regr.rpart cv 1
#> 4: 0 easy regr.rpart cv 1
#> 5: 0 easy regr.rpart cv 1
#> 6: 0 easy regr.rpart cv 1
#> 7: 0 easy regr.rpart cv 1
#> 8: 0 easy regr.rpart cv 1
#> 9: 0 easy regr.rpart cv 1
#> 10: 0 impossible regr.rpart cv 1
#> 11: 0 impossible regr.rpart cv 1
#> 12: 0 impossible regr.rpart cv 1
#> 13: 0 impossible regr.rpart cv 1
#> 14: 0 impossible regr.rpart cv 1
#>
#> $results.csv
#> grid_job_i task.i learner.i resampling.i task_id learner_id
#> <int> <int> <int> <int> <char> <char>
#> 1: 1 1 1 1 easy regr.featureless
#> 2: 2 1 2 1 easy regr.rpart
#> 3: 3 2 1 1 impossible regr.featureless
#> 4: 4 2 2 1 impossible regr.rpart
#> resampling_id iteration start.time end.time process
#> <char> <int> <POSc> <POSc> <int>
#> 1: cv 1 2026-04-28 14:59:38 2026-04-28 14:59:38 26324
#> 2: cv 1 2026-04-28 14:59:38 2026-04-28 14:59:38 26324
#> 3: cv 1 2026-04-28 14:59:38 2026-04-28 14:59:38 26324
#> 4: cv 1 2026-04-28 14:59:38 2026-04-28 14:59:38 26324
#> regr.rmse regr.mae
#> <num> <num>
#> 1: 1.3937502 1.1580176
#> 2: 0.6459229 0.5284654
#> 3: 1.7017688 1.3413757
#> 4: 1.5258893 0.9700115
11 P-values for comparing Same/Other/All training
Description
Same/Other/All K-fold cross-validation (SOAK) results in K measures of test error/accuracy. This function computes P-values (two-sided T-test) between Same and All/Other.
Usage
pvalue(score_in, value.var = NULL, digits=3)Arguments
score_in |
Data table output from |
value.var |
Name of column to use as the evaluation metric in T-test. Default
NULL means to use the first column matching |
digits |
Number of decimal places to show for mean and standard deviation. |
Value
List of class "pvalue" with named elements value.var,
stats, pvalues.
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=rep(1:2, each=0.5*N))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^person)
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
yname <- paste0("y_",pattern)
reg.dt[, (yname) := f(x,person)+rnorm(N, sd=0.5)][]
task.dt <- reg.dt[, c("x","person",yname), with=FALSE]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target=yname)
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
(bench.grid <- mlr3::benchmark_grid(
reg.task.list,
reg.learner.list,
SOAK))
#> task learner resampling
#> <char> <char> <char>
#> 1: easy regr.featureless same_other_sizes_cv
#> 2: easy regr.rpart same_other_sizes_cv
#> 3: impossible regr.featureless same_other_sizes_cv
#> 4: impossible regr.rpart same_other_sizes_cv
bench.result <- mlr3::benchmark(bench.grid)
#> INFO [14:59:39.061] [mlr3] Running benchmark with 72 resampling iterations
#> INFO [14:59:39.117] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 1/18)
#> INFO [14:59:39.134] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 2/18)
#> INFO [14:59:39.145] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 3/18)
#> INFO [14:59:39.155] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 4/18)
#> INFO [14:59:39.166] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 5/18)
#> INFO [14:59:39.176] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 6/18)
#> INFO [14:59:39.187] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 7/18)
#> INFO [14:59:39.198] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 8/18)
#> INFO [14:59:39.215] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 9/18)
#> INFO [14:59:39.226] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 10/18)
#> INFO [14:59:39.236] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 11/18)
#> INFO [14:59:39.247] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 12/18)
#> INFO [14:59:39.257] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 13/18)
#> INFO [14:59:39.268] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 14/18)
#> INFO [14:59:39.278] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 15/18)
#> INFO [14:59:39.288] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 16/18)
#> INFO [14:59:39.299] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 17/18)
#> INFO [14:59:39.309] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 18/18)
#> INFO [14:59:39.320] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 1/18)
#> INFO [14:59:39.334] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 2/18)
#> INFO [14:59:39.347] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 3/18)
#> INFO [14:59:39.361] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 4/18)
#> INFO [14:59:39.374] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 5/18)
#> INFO [14:59:39.388] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 6/18)
#> INFO [14:59:39.401] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 7/18)
#> INFO [14:59:39.415] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 8/18)
#> INFO [14:59:39.429] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 9/18)
#> INFO [14:59:39.448] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 10/18)
#> INFO [14:59:39.462] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 11/18)
#> INFO [14:59:39.475] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 12/18)
#> INFO [14:59:39.488] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 13/18)
#> INFO [14:59:39.502] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 14/18)
#> INFO [14:59:39.515] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 15/18)
#> INFO [14:59:39.529] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 16/18)
#> INFO [14:59:39.542] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 17/18)
#> INFO [14:59:39.556] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 18/18)
#> INFO [14:59:39.569] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 1/18)
#> INFO [14:59:39.580] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 2/18)
#> INFO [14:59:39.591] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 3/18)
#> INFO [14:59:39.601] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 4/18)
#> INFO [14:59:39.612] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 5/18)
#> INFO [14:59:39.622] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 6/18)
#> INFO [14:59:39.633] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 7/18)
#> INFO [14:59:39.643] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 8/18)
#> INFO [14:59:39.660] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 9/18)
#> INFO [14:59:39.670] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 10/18)
#> INFO [14:59:39.681] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 11/18)
#> INFO [14:59:39.691] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 12/18)
#> INFO [14:59:39.702] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 13/18)
#> INFO [14:59:39.712] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 14/18)
#> INFO [14:59:39.723] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 15/18)
#> INFO [14:59:39.733] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 16/18)
#> INFO [14:59:39.744] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 17/18)
#> INFO [14:59:39.754] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 18/18)
#> INFO [14:59:39.765] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 1/18)
#> INFO [14:59:39.779] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 2/18)
#> INFO [14:59:39.793] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 3/18)
#> INFO [14:59:39.806] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 4/18)
#> INFO [14:59:39.820] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 5/18)
#> INFO [14:59:39.833] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 6/18)
#> INFO [14:59:39.847] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 7/18)
#> INFO [14:59:39.860] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 8/18)
#> INFO [14:59:39.880] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 9/18)
#> INFO [14:59:39.893] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 10/18)
#> INFO [14:59:39.907] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 11/18)
#> INFO [14:59:39.920] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 12/18)
#> INFO [14:59:39.933] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 13/18)
#> INFO [14:59:39.947] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 14/18)
#> INFO [14:59:39.960] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 15/18)
#> INFO [14:59:39.974] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 16/18)
#> INFO [14:59:39.987] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 17/18)
#> INFO [14:59:40.001] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 18/18)
#> INFO [14:59:40.028] [mlr3] Finished benchmark
bench.score <- mlr3resampling::score(bench.result, mlr3::msr("regr.rmse"))
bench.plist <- mlr3resampling::pvalue(bench.score)
plot(bench.plist)
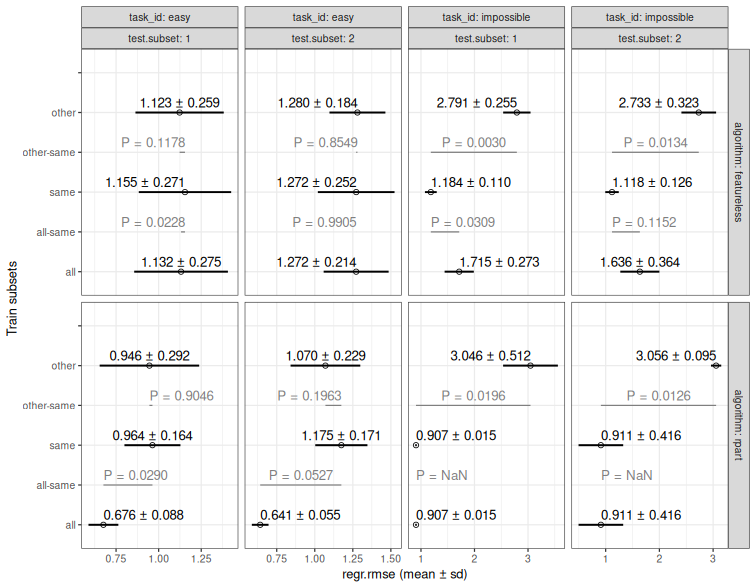
12 P-values for full versus down-sampled SOAK results
Description
For one test.subset, compute summary statistics and paired
two-sided t-test P-values that compare same versus
all/other, for two train-size panels:
full (n.train.groups == groups) and down-sampled
(n.train.groups == min(groups)).
Usage
pvalue_downsample(
score_in,
value.var = NULL,
digits = 3
)Arguments
score_in |
Data table output from |
value.var |
Name of column to use as the evaluation metric in T-test. Default
NULL means to use the first column matching |
digits |
Non-negative integer number of digits to print in mean/SD text labels. |
Value
List of class "pvalue_downsample" with named elements
subset_name, model_name, value.var,
stats, and pvalues. subset_name and
model_name are inferred from score_in.
Author(s)
Daniel Agyapong
Examples
library(data.table)
set.seed(1)
N <- 120L
x <- runif(N, -3, 3)
sim.dt <- data.table(
x = x,
y = sin(x) + rnorm(N, sd = 0.3),
Subset = rep(c("large", "large", "small"), length.out = N)
)
task <- mlr3::TaskRegr$new("iid_easy", sim.dt, target = "y")
task$col_roles$feature <- "x"
task$col_roles$subset <- "Subset"
soak <- mlr3resampling::ResamplingSameOtherSizesCV$new()
soak$param_set$values$folds <- 2L
soak$param_set$values$seeds <- 1L
soak$param_set$values$sizes <- 0L
if(requireNamespace("rpart", quietly = TRUE)){
grid <- mlr3::benchmark_grid(task, list(mlr3::lrn("regr.rpart")), soak)
result <- mlr3::benchmark(grid)
score.dt <- mlr3resampling::score(result)[, .(
task_id, test.subset, test.fold, train.subsets,
groups, n.train.groups, algorithm, RMSE = sqrt(regr.mse)
)]
down.list <- mlr3resampling::pvalue_downsample(
score.dt[task_id == "iid_easy" & test.subset == "small" & algorithm == "rpart"],
value.var = "RMSE",
digits = 3
)
if(requireNamespace("ggplot2", quietly = TRUE)) plot(down.list)
}
#> INFO [14:59:41.485] [mlr3] Running benchmark with 20 resampling iterations
#> INFO [14:59:41.503] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 1/20)
#> INFO [14:59:41.518] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 2/20)
#> INFO [14:59:41.532] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 3/20)
#> INFO [14:59:41.547] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 4/20)
#> INFO [14:59:41.569] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 5/20)
#> INFO [14:59:41.585] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 6/20)
#> INFO [14:59:41.599] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 7/20)
#> INFO [14:59:41.613] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 8/20)
#> INFO [14:59:41.626] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 9/20)
#> INFO [14:59:41.640] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 10/20)
#> INFO [14:59:41.654] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 11/20)
#> INFO [14:59:41.668] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 12/20)
#> INFO [14:59:41.682] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 13/20)
#> INFO [14:59:41.696] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 14/20)
#> INFO [14:59:41.710] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 15/20)
#> INFO [14:59:41.733] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 16/20)
#> INFO [14:59:41.747] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 17/20)
#> INFO [14:59:41.760] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 18/20)
#> INFO [14:59:41.774] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 19/20)
#> INFO [14:59:41.788] [mlr3] Applying learner 'regr.rpart' on task 'iid_easy' (iter 20/20)
#> INFO [14:59:41.804] [mlr3] Finished benchmark
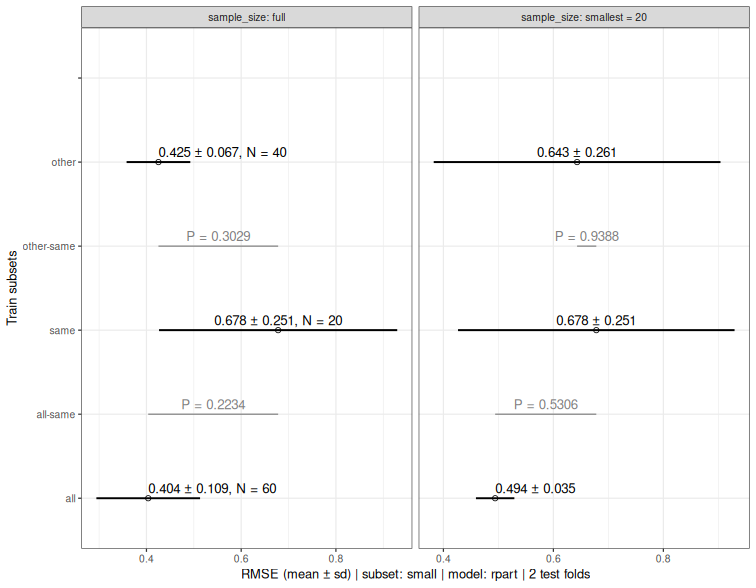
13 Score benchmark results
Description
Computes a data table of scores.
Usage
score(bench.result, ...)Arguments
bench.result |
Output of |
... |
Additional arguments to pass to |
Value
data table with scores.
Author(s)
Toby Dylan Hocking
Examples
N <- 80
library(data.table)
set.seed(1)
reg.dt <- data.table(
x=runif(N, -2, 2),
person=rep(1:2, each=0.5*N))
reg.pattern.list <- list(
easy=function(x, person)x^2,
impossible=function(x, person)(x^2)*(-1)^person)
SOAK <- mlr3resampling::ResamplingSameOtherSizesCV$new()
reg.task.list <- list()
for(pattern in names(reg.pattern.list)){
f <- reg.pattern.list[[pattern]]
yname <- paste0("y_",pattern)
reg.dt[, (yname) := f(x,person)+rnorm(N, sd=0.5)][]
task.dt <- reg.dt[, c("x","person",yname), with=FALSE]
task.obj <- mlr3::TaskRegr$new(
pattern, task.dt, target=yname)
task.obj$col_roles$stratum <- "person"
task.obj$col_roles$subset <- "person"
reg.task.list[[pattern]] <- task.obj
}
reg.learner.list <- list(
mlr3::LearnerRegrFeatureless$new())
if(requireNamespace("rpart")){
reg.learner.list$rpart <- mlr3::LearnerRegrRpart$new()
}
(bench.grid <- mlr3::benchmark_grid(
reg.task.list,
reg.learner.list,
SOAK))
#> task learner resampling
#> <char> <char> <char>
#> 1: easy regr.featureless same_other_sizes_cv
#> 2: easy regr.rpart same_other_sizes_cv
#> 3: impossible regr.featureless same_other_sizes_cv
#> 4: impossible regr.rpart same_other_sizes_cv
bench.result <- mlr3::benchmark(bench.grid)
#> INFO [14:59:42.490] [mlr3] Running benchmark with 72 resampling iterations
#> INFO [14:59:42.499] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 1/18)
#> INFO [14:59:42.510] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 2/18)
#> INFO [14:59:42.522] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 3/18)
#> INFO [14:59:42.533] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 4/18)
#> INFO [14:59:42.544] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 5/18)
#> INFO [14:59:42.555] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 6/18)
#> INFO [14:59:42.566] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 7/18)
#> INFO [14:59:42.586] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 8/18)
#> INFO [14:59:42.598] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 9/18)
#> INFO [14:59:42.608] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 10/18)
#> INFO [14:59:42.620] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 11/18)
#> INFO [14:59:42.631] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 12/18)
#> INFO [14:59:42.642] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 13/18)
#> INFO [14:59:42.653] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 14/18)
#> INFO [14:59:42.664] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 15/18)
#> INFO [14:59:42.676] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 16/18)
#> INFO [14:59:42.687] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 17/18)
#> INFO [14:59:42.698] [mlr3] Applying learner 'regr.featureless' on task 'easy' (iter 18/18)
#> INFO [14:59:42.709] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 1/18)
#> INFO [14:59:42.732] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 2/18)
#> INFO [14:59:42.748] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 3/18)
#> INFO [14:59:42.762] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 4/18)
#> INFO [14:59:42.776] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 5/18)
#> INFO [14:59:42.790] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 6/18)
#> INFO [14:59:42.804] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 7/18)
#> INFO [14:59:42.818] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 8/18)
#> INFO [14:59:42.832] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 9/18)
#> INFO [14:59:42.846] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 10/18)
#> INFO [14:59:42.860] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 11/18)
#> INFO [14:59:42.884] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 12/18)
#> INFO [14:59:42.899] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 13/18)
#> INFO [14:59:42.913] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 14/18)
#> INFO [14:59:42.926] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 15/18)
#> INFO [14:59:42.940] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 16/18)
#> INFO [14:59:42.954] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 17/18)
#> INFO [14:59:42.969] [mlr3] Applying learner 'regr.rpart' on task 'easy' (iter 18/18)
#> INFO [14:59:42.983] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 1/18)
#> INFO [14:59:42.994] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 2/18)
#> INFO [14:59:43.005] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 3/18)
#> INFO [14:59:43.025] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 4/18)
#> INFO [14:59:43.038] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 5/18)
#> INFO [14:59:43.049] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 6/18)
#> INFO [14:59:43.059] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 7/18)
#> INFO [14:59:43.070] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 8/18)
#> INFO [14:59:43.081] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 9/18)
#> INFO [14:59:43.092] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 10/18)
#> INFO [14:59:43.103] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 11/18)
#> INFO [14:59:43.115] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 12/18)
#> INFO [14:59:43.126] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 13/18)
#> INFO [14:59:43.137] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 14/18)
#> INFO [14:59:43.148] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 15/18)
#> INFO [14:59:43.166] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 16/18)
#> INFO [14:59:43.180] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 17/18)
#> INFO [14:59:43.191] [mlr3] Applying learner 'regr.featureless' on task 'impossible' (iter 18/18)
#> INFO [14:59:43.201] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 1/18)
#> INFO [14:59:43.216] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 2/18)
#> INFO [14:59:43.230] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 3/18)
#> INFO [14:59:43.247] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 4/18)
#> INFO [14:59:43.261] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 5/18)
#> INFO [14:59:43.275] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 6/18)
#> INFO [14:59:43.290] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 7/18)
#> INFO [14:59:43.310] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 8/18)
#> INFO [14:59:43.328] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 9/18)
#> INFO [14:59:43.342] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 10/18)
#> INFO [14:59:43.356] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 11/18)
#> INFO [14:59:43.370] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 12/18)
#> INFO [14:59:43.384] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 13/18)
#> INFO [14:59:43.398] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 14/18)
#> INFO [14:59:43.412] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 15/18)
#> INFO [14:59:43.426] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 16/18)
#> INFO [14:59:43.440] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 17/18)
#> INFO [14:59:43.462] [mlr3] Applying learner 'regr.rpart' on task 'impossible' (iter 18/18)
#> INFO [14:59:43.491] [mlr3] Finished benchmark
bench.score <- mlr3resampling::score(bench.result, mlr3::msr("regr.rmse"))
plot(bench.score)